Summer Research Program 2026
Biology || Chemistry || Computer Science and Software Engineering || Mathematics
Biology
Project: eDNA Biodiversity in Marine Environments
Faculty Mentor: Jason E. Adolf, Ph.D.
Biodiversity is an important ecological measurement used to assess marine communities. Traditionally this is done by catching, identifying and counting organisms, whether ‘micro’ or ‘macro’. While capture surveys are still important, genomics-based technologies are being developed that allow biodiversity of a wide range of organisms to be determined by sequencing DNA left behind in the environment where they live, so called environmental DNA or ‘eDNA’. Projects this summer will focus on using eDNA to determine fish, invertebrate and prokaryotic biodiversity in regional waters. Projects will involve a combination of field sampling, laboratory analyses, as well as bioinformatic and ecological analyses on the computer, and preparation of results for professional presentation.
Project: Climate Change Impacts on Maritime Forests
Faculty Mentor: Pedram Daneshgar, Ph.D.
Maritime forests are rare and threatened ecosystems that occur in close proximity to the ocean where they are constantly exposed to salt. Because of their location they are often threatened by development, but also by climate change impacts such a rising sea level. The Daneshgar lab will be exploring the impacts of sea level rise and salt marsh encroachment on juvenile maritime forest species through greenhouse experiments. Students will also participate in outreach efforts to educate communities about climate impacts on maritime forests.
Project: Sustainable Agriculture and Health
Faculty Mentor: Pedram Daneshgar, Ph.D.
The productivity and health benefits of many of our important agricultural crops are dependent on environment and sustainable practices. Through field and greenhouse experiments, the Daneshgar lab will explore how environmental stress and agricultural practice may impact the productivity of two major crops: coffee and blueberries. To follow, through lab work, the group examine how environment shapes the health benefits offered by these crops.
Project: Stress Signaling in Normal and Cancerous Cells
Faculty Mentors: Dottie Lobo, Ph.D., and James Mack, Ed.D
One project to be addressed is the influence of specific essential oils on cell signaling pathways and the stress response of normal and cancerous cells grown in culture. With the rise of antibiotic resistance, there has been increased emphasis on alternative, natural products that may be used in medicine. Previous research in this laboratory has indicated that cypress essential oil treatment leads to decreased proliferation and apoptosis in a variety of human cell lines. This summer, we would like to continue to study the effects of cypress essential oils on the control of proliferation and apoptosis in cancer cell lines, and to compare all results to normal cell lines. This project should be nearing completion, and work on additional essential oils may be initiated. A second project may be done to evaluate the role of hypoxia in stress signaling. Additionally, there has been work performed to characterize the anti-bacterial role of essential oils at Monmouth, We will also address the effects of specific essential oils on the growth of multidrug resistant bacteria including: Acinobacter baumanni, Enterobacter cloacae, and Klebsiella pneumonia.
Project: Genetic and Physiological Regulation of Aggression Using Drosophila Melanogaster as a Model
Faculty Mentor: Saheli Sengupta, Ph.D.
Aggression is a complex and evolutionarily conserved behavior shaped by genetic, neural, and physiological states. Dysregulation of aggression is associated with multiple neurological and neurodegenerative disorders, yet the mechanisms by which molecular perturbations and internal states influence aggressive behavior remain poorly understood. This project uses Drosophila melanogaster as a genetically tractable model to investigate how defined genetic insults and metabolic states regulate aggression and related behaviors.
Data generated from these two projects will contribute to peer-reviewed publications and support future external grant submissions.
Project 1
Project 1 focuses on characterizing the behavioral consequences of expressing full-length human Ataxin-2 (ATXN2) containing an expanded polyglutamine tract (117Q), a protein associated with neurodegenerative disease. Using the UAS/GAL4 system, ATXN2-117Q has been expressed in select neuronal populations in the fly brain, including the mushroom bodies, which play key roles in learning and memory. Students will assess whether expression of this protein is toxic to specific neuronal populations by systematically quantifying changes in aggression, courtship, and locomotion. Behavioral analyses will be paired with anatomical and neurochemical characterization of affected neurons. Identification of the neurochemical identity of vulnerable neuronal populations will be conducted in collaboration with McLean Hospital, Harvard Medical School. These experiments will enable students to link neuronal vulnerability to specific behavioral outcomes and identify candidate neuronal populations and molecular pathways for future targeted genetic and functional studies.
Project 2
Project 2 examines the relationship between hunger and aggression. Students will establish a starvation paradigm that restricts food while preventing dehydration and assess how different durations of starvation (24, 48, and 72 hours) alter aggressive behavior in both male and female flies. To uncover molecular mechanisms underlying hunger-induced changes in aggression, bulk RNA sequencing will be performed on fly brains across starvation conditions. Differential gene expression analysis will be used to identify pathways and candidate genes associated with altered aggressive states, providing a foundation for follow-up functional experiments.
Project: Songbird and Drone Ecology in Suburbia
Faculty Mentor: Sean Sterrett, Ph.D.
Wildlife populations are constantly under pressure from local, national and global declines due to human activity. Urbanization and suburbanization are types of development that create challenging stressors for animals to persist (i.e., habitat loss, increasing levels of disease, collection for pet trade). These two projects aim to focus on wildlife populations in suburban environments, but from contrasting viewpoints; using drones to study turtle populations and evaluating the impacts of sound pollution on a highly vocal and diverse songbirds across New Jersey.
There is growing evidence that drones have the potential to improve sampling for aquatic and terrestrial wildlife. However, there remains a need to evaluate the efficacy of these novel methods relative to traditional sampling protocols. As part of an external grant on developing drone-based protocols for sampling Diamond-backed terrapin (DT) populations, we will take the opportunity to collect drone images of DT populations and compare these to more traditional approaches for sampling DT populations (i.e., head count surveys).
Human-mediated impacts to forested ecosystems are clearly on the rise and these threats are often increased in suburban areas. Songbirds (suborder Passeri) are a highly diverse taxon in New Jersey with its proximity to the Atlantic Flyway, a major bird migration route. As a result of significant human development, the landscape of sound in New Jersey forests has changed (i.e., increased, chronic). Songbirds seasonally communicate vocally to attract mates, defend territories and warn of predators. Due to the change in landscape, we want to evaluate how songbird communities are influenced by sound pollution. We will approach this objective by determining representative areas with common songbird communities that have varying levels of ambient anthropogenic sound using Wildlife Acoustics Song Meter SM4 Acoustic Recorders and develop a sampling design for testing the hypothesis related to community metrics.
Project: Genetic Analysis of a Novel Chromatin Assembly Mutant in Drosophila
Faculty Mentor: Jennifer Urban, Ph.D.
My lab is interested in how DNA replication-coupled chromatin assembly influences cell fate decisions during asymmetric cell division (ACD) in the Drosophila germline. ACD contributes to cellular diversity by producing two genetically identical daughter cells that adopt different cell fates. Disruption of this process can lead to infertility, tissue degeneration, and cancer. Histone chaperone proteins are known to regulation chromatin reassembly during DNA replication, however how these factors influence cell fate specification in a multicellular organism remains poorly understood.
To address this question, we will use Drosophila genetics to characterize a novel mutant allele I generated in a gene known to function in replication-coupled chromatin assembly. Preliminary observations revealed difficulty recovering adult males carrying the mutation, suggesting potential defects in viability or germline development. Two summer students will first determine whether mutant animals survive at expected Mendelian ratios, testing for developmental lethality. We will then assess reproductive fitness by measuring male and female fertility. Together these approaches will determine whether the mutation disrupts viability, germline development, or fertility. This work will provide foundational insight into how DNA replication machinery contributes to stem cell maintenance in vivo. In addition to generating critical preliminary data for future mechanistic studies, the project will offer students hands-on training in classical genetics, experimental design, and data analysis.
Chemistry
Project: Exploring Local Structures in DNA Conformational Transitions with Fluorescent Base Analogs
Faculty Mentor: Davis Jose, Ph.D.
Investigate how local base-stacking and hydrogen-bonding rearrangements facilitate the interconversion between various DNA conformations, and examine how small-molecule ligands influence these transitions at specific nucleotide sites.
Background and Significance
DNA, the carrier of genetic information in living organisms, is highly adaptable and exhibits conformational polymorphism. Besides the well-known Watson-Crick B-form double helix, it can also assume structures like A-form, Z-form, G-quadruplexes (GQ), and i-motif. This polymorphism is essential for many key biological functions. For example, A-form DNA facilitates DNA packaging, while G-quadruplex structures in guanine-rich regions of DNA and RNA are important in cancer and aging. DNA’s dynamic nature allows it to switch between conformations easily, which has diverse applications. However, the exact mechanisms of DNA conformational interconversion remain experimentally unexplored. Gaining this understanding is crucial for unlocking DNA’s potential in biotechnology.
Methods and Approaches
Optical spectroscopy methods with fluorescent and CD-active base analogues, site-specifically placed into nucleic acid frameworks, will monitor global and local conformational changes. This unique approach tracks individual base-pair changes at specific locations, without interference from canonical bases, revealing base-pair orientation, stacking perturbations, and backbone fluctuations at single-base-pair resolution during transitions. Synthetic small organic molecules will be introduced to dissect their contributions. Results from multiple techniques will be integrated to map basic mechanisms and pathways.
Expected Outcomes
This research aims to clarify how the local DNA conformation changes during various transitions, thereby improving understanding of processes such as gene regulation, telomere maintenance, cancer, and aging. These insights will guide drug development, such as GQ stabilizers to prevent tumor growth, or the identification of disease origins, such as GQ unwinding defects in premature aging syndromes.
Project: Substituted Quinoline Complexes of the Group 6 Transition Metals
Faculty Mentor: Gregory Moehring, Ph.D.
Quinoline-8-carbaldehyde (Q8C, Figure 1) is a substance of considerable interest. Q8C serves as a substrate for several catalytic transformations. Q8C, or its derivative quinoline-8-acyl (Q8acyl) also serves as a ligand (a supporting portion) for molecular substances built around later transition metals such as rhodium, iridium, platinum, and palladium. Our research group recently expanded the chemistry of Q8C and its derivatives by binding Q8C and its Q8acyl derivative to rhenium, a mid-transition metal. We are interested in reactions of Q8C with rhenium-containing complexes for two reasons: 1) Q8C provided an excellent candidate substrate for the exploration of a new rhenium-based C-H bond activation and 2) Q8C and its derivatives, when bound to rhenium, can provide a potential scaffold on which to build new rhenium-centered cytotoxins (potential chemotherapeutics) or cell imaging agents (substances that are useful in examining mechanisms of cytotoxicity). Our group has an interest in rhenium-based cytotoxins.
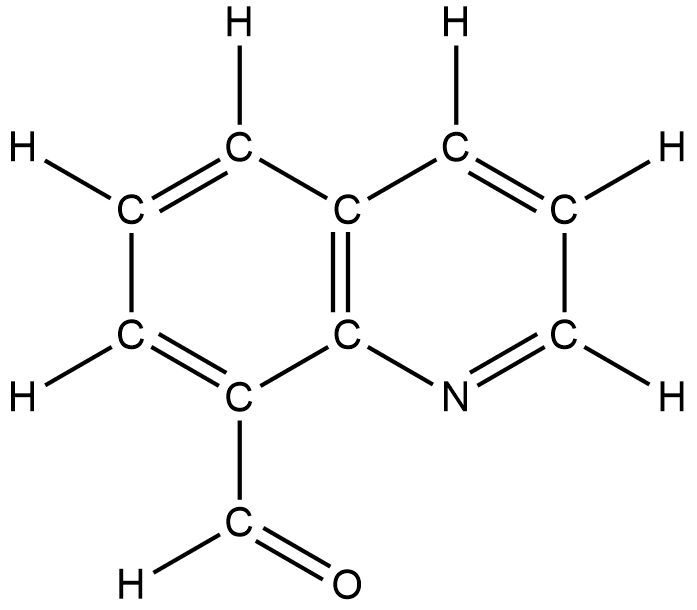
This summer, we look to prepare and characterize Q8C-supported complexes of chromium, molybdenum, and tungsten (the Group 6 metals of the Periodic Table). The proposed research continues our search for inorganic-based cytotoxins as potential chemotherapeutics. Certain molybdenum-containing complexes with two or more supporting carbonyl groups are cytotoxic. Furthermore, our work with rhenium has shown a remarkably low energy barrier for the substitution of bidentate Q8C ligands by monodentate ligands such as acetonitrile or dimethyl sulfoxide. We plan to use the series of M(CO)4(Q8C) (M = Cr, Mo, or W) target substances to further explore the substitution of transition metal-bound bidentate Q8C ligands by monodentate ligands. Specifically, we want to determine if the formally zero oxidation state target substances, M(CO)4(Q8C), make the Q8C ligand even more readily substituted than in the comparable rhenium(I)-based ReBr(CO)3(Q8C) complex.
Project: Structure/Function of Nucleic Acids: Aptamer, Riboswitch and DNA Enzyme
Faculty Mentor: Jonathan Ouellet, Ph.D.
Research Question or Problem
The lab is interested in synthetic biology, where the nucleic acids can specifically bind a small molecule to relate an action. This summer’s projects further the overall lab goals. Among the projects, one is to test the efficiency of a fluorescence RNA aptamer by adding new chemical compounds that can interfere with the RNA G-Quadruplex structure. Another project is to perform fluorescence kinetics of a DNA enzyme that cuts DNA in presence of Zn++.
Background and Significance
Several RNA G-Quadruplexes have been found in viruses and some mammalian genes. In viruses, the G-Quadruplex seems to be involved in the reproduction live cycle of the virus. Interfering with that timing was shown to lower virus titers. Since the fluorescence of the RNA aptamer is directly linked to the structure of the G-Quadruplex, it is used as a signal to indicate the stability of the G-Quadruplex in presence of new compounds.
The existence of a DNA enzyme cutting DNA is extremely rare. Very little is known about hat DNA enzyme. The first step in understanding its cleavage mechanism for optimization is to understand its kinetics. Using a DNA substrate containing 2-aminopurine, the kinetic can be measured using fluorescence spectroscopy.
Methods and Approaches
The students will perform PCR and in vitro transcription to produce RNA. After acrylamide gel purification, the RNA is used with fluorescence spectroscopy by performing fluorescence melting curves. The extracted Tm (melting temperature) is an indication of the G-Quadruplex stability. The melting curves are then measured again, but in the presence of the various new chemical compounds.
The DNA Enzyme and fluorescent Substrate are incubated in their buffer solution in a thermoregulated cuvette into the fluorescence spectrometer. Last year, we purchased a special spectrophotometer lid that accepts a needle to inject sample into the cuvette through a small hole. Here, Zn++ will be injected to initiate the cleavage catalysis. The real-time monitoring of fluorescence will allow to calculate the observed rate of cleavage of the reaction.
Project: Separation of Anions Using Metal Organic Frameworks
Faculty Mentor: Debmalya Ray, Ph.D.
Developing energy-efficient separations can save the US petrochemical sectors 100 million tons in CO2 emission and US $4 billion in energy costs annually. Most commonly used separation process (i.e., distillation) accounts for 10–15% of the world’s energy consumption. Thus, development of inexpensive and energy-efficient separations techniques based on chemical properties or size are the need of the century.
Metal-organic frameworks (MOFs) are a class of highly crystalline, porous, stable, and extremely tunable materials with limitless designing space. This makes MOFs ideal candidate for separation applications. Thus, in this research direction, I plan to study separation of anions (NO3–, TcO4–, ReO4–, HMoO4–, SO42-, BiO3–). Separation of ions like TcO4– from other anions is important for nuclear waste management purpose as the high mobility and volatility of TcO4– ion is one of the major safety challenges, relevant to the nuclear industry. Thus, in this project we will design metal organic frameworks which can effectively separate TcO4– from other anions mentioned above. We will use density functional theory and ab-initio molecular dynamics-based approach to study such chemical separation processes. We expect to publish our result in American Chemical Society journals and present them in national and local conferences.
Project: Surface Aging and Mineral Coating of Microplastics in Marine Environments: Implications for Contaminant Mobility
Faculty Mentor: Tsanangurayi Tongesayi, Ph.D.
Microplastics (MPs) in marine environments undergo physical, chemical, and biological aging that alters their surfaces and affects how they interact with contaminants. This interdisciplinary project investigates how MPs—of various polymer types collected from coastal waters—change through exposure to minerals, natural organic matter, sunlight, and environmental conditions. The work integrates analytical chemistry, marine sampling, environmental science, and materials characterization to understand how these transformations influence contaminant mobility.
Preliminary data from my research lab provides a foundation for this work. Laboratory experiments using polyethylene terephthalate microplastics (PET‑MP) show that minerals such as goethite (GT) and kaolin (KL) attach to MP surfaces through both physical and chemical mechanisms. FT‑IR analysis reveals shifts in O–H stretching vibrations, decreases in PET ester functional group intensities, and spectral broadening, indicating hydrogen bonding and van der Waals interactions for KL, and hydrogen bonding plus electrostatic interactions for GT. XRF confirms substantial GT adsorption (Fe ≈ 9190 ± 50 ppm), while XPS shows shifts in C1s and O1s binding energies, demonstrating changes in surface functional groups and mineral attachment. GT forms localized reactive patches, whereas KL produces broader coatings.
Students will compare these laboratory‑aged MPs with marine‑aged MPs collected from local waters. Because environmental MPs experience much longer and more variable aging times, comparisons will explicitly account for differences in exposure duration, environmental conditions, and polymer type. Students will identify and characterize marine MPs before comparing their surface chemistry to laboratory‑aged samples.
The project will address three interdisciplinary questions:
- How do these changes affect contaminant mobility, microplastic sedimentation, and biogeochemical cycling?
- How do marine and laboratory aging processes modify MP surface chemistry and structure?
- How do mineral and organic coatings influence adsorption of metals and metalloids?
Computer Science and Software Engineering
Project: Quantum Computing for Cybersecurity Applications
Faculty Mentor: Brian Callahan, Ph.D.
Quantum computing is a new computing paradigm that is beginning to show promise in a wide variety of fields such as drug discovery and finance. Able to mine through high-dimensionality data in ways that will be forever impossible to classical computers, now is the perfect time to start exploiting quantum’s power in other fields with data of similar shapes. Cybersecurity is such a field, and one where quantum computing is thought of as a negative due to quantum’s potential for disruption in cracking cryptography, and, therefore, stealing secrets.
This project seeks to continue already published work by the faculty sponsor (Cotrupi and Callahan 2025), grant-funded research by current MU students (Ramirez Velandia, Cotrupi, and Callahan forthcoming), and other work by Monmouth University Cybersecurity Research Center students (Carducci forthcoming). This research lays the groundwork to build a commercial-off-the-shelf product for cybersecurity professionals that harnesses the power of quantum computing to accelerate the discovery of Indicators of Compromise, including malicious network traffic, phishing emails, and anomalous user behavior. We aim to revolutionize how organizations protect themselves against modern cyber threats by providing them with a platform that will deliver speed and accuracy that can never be matched by classical machines.
This project will engage in cutting-edge quantum algorithm and quantum machine learning technique development, incorporating current state-of-the-art techniques such as QSVM and quantum kernels, and developing optimizations for these techniques on a real-world quantum computer, an IBM Quantum System One. We may potentially develop new quantum algorithms and quantum machine learning techniques. Programming will be done using IBM’s Qiskit IDE, eventually being made accessible through a web application.
Immediate expected outcomes are journal articles and conference presentations. Long-term expected outcomes are patents and potentially a commercial entity to sell and support our quantum cybersecurity platform to companies worldwide.
Project: Implementing GArel: Bidirectional Type Checking for Relational Array Analysis
Faculty Mentor: Weihao Qu, Ph.D.
This research focuses on the practical implementation of GArel, a relational type and effect system designed to verify quantitative properties in programs that manipulate mutable arrays. While theoretical frameworks for relational cost analysis exist, their implementation often suffers from high complexity and low automation. To address these challenges, this project focuses on developing a high-performance type checker utilizing bidirectional type checking techniques.
Bidirectional typing is a powerful algorithmic strategy that splits type checking into two modes—inference and checking—greatly reducing the need for manual type annotations and improving error reporting. By applying this technique to GArel, we can more efficiently track customizable relations within array information and manage the system’s parameterizable grading mechanism. This implementation is critical for scaling GArel to handle complex, real-world functional programs that rely on mutable state.
The student researcher will work on translating GArel’s theoretical rules into an algorithmic formulation suitable for bidirectional checking. This involves integrating the checker with advanced SMT solvers to automate the verification of quantitative relational properties, such as relative execution cost or memory usage. The resulting tool will serve as a practical platform for reasoning about program behavior, directly contributing to the state of the art in formal program analysis. This work is part of a larger research initiative supported by an NSF CRII grant (#2451348) focused on formal verification.
Mathematics
Project: A Mathematical Model of Misinformation
Faculty Mentor: Torrey Gallagher, Ph.D.
This project will study how the opinions of individuals in groups evolve over time, and the ways in which individuals can strategically influence the dynamics of a group. This problem has classical roots in the mathematical economics fields of Decision Theory and Social Choice Theory and was first studied by Morris DeGroot who proposed a model in which trust between individuals remained constant throughout time, and opinion updates occurred in discrete time intervals. This model has many interesting features and leads to useful and quantifiable long-term dynamics (such as the group reaching a consensus versus forming multiple subgroups with differing opinions versus periodic cycling of opinions). However, it has some significantly unrealistic assumptions; most notably, the assumption that trust between individuals remains fixed throughout time.
In this project, we will consider reasonable criteria that should govern trust updates and study the resulting long-term dynamics. Of particular interest will be classifying the initial opinions which lead to consensus versus polarization, and the effect of agents supplying misinformation on the long-term group dynamics. The problem sits naturally at the intersection of graph theory, matrix analysis (linear algebra), and numerical analysis. As such, a mixture of pen-and-paper analysis and coding experiments will be required.
