2026 Student Research Conference Abstracts
24th Annual School of Science Student Research Conference
April 10, 2026
Edison 201 (Science Building Multipurpose Room)
Featuring Poster Presentations of Student Research
Agenda
| Time | Event |
|---|---|
| 10–11 a.m. | Registration and Poster Set-up |
| 11–11:15 a.m. | Welcome and Opening Remarks Dean Joe Coyle |
| 11:15 a.m.–1:15 p.m. | Poster Presentations of Student Research from the School of Science |
| 1:15–1:45 p.m. | Closing Remarks and Presentation of the Dean’s Awards for Excellence in Undergraduate Research Dean Joe Coyle |
Abstracts
- Agenda
- Department of Biology
- Department of Chemsitry and Physics
- Department of Computer Science and Software Engineering
- Department of Mathematics
Department of Biology
BY-1: Disturbance Ecology and Divergent Recovery of the Atlantic Mole Crab
Jessica Kipnis and Timothy Buckley (Department of Biology)
Faculty Mentors: Jason Adolf, Ph.D., and Richard Bastian, Ph.D.
Read the Abstract—BY-1: Disturbance Ecology and Divergent Recovery of the Atlantic Mole Crab
Sandy beach ecosystems are increasingly impacted by coastal engineering and climate variability, yet the demographic mechanisms driving bioindicator population responses to disturbance remain poorly understood. This study examined population dynamics of the Atlantic mole crab (Emerita talpoida) across Monmouth County from 2021-2025, spanning baseline conditions, active disturbance, and post-disturbance recovery. Mole crabs were sampled across multiple sites and surf zone habitats (bore and swash), and categorized into three size classes representing juvenile, subadult, and adult cohorts. Using negative binomial generalized linear mixed models, we quantified the effects of year, microhabitat, and water temperature on abundance while accounting for sampling effort and sampling site differences. Population structure remained stable during baseline years (2021-2022), but a significant demographic shift occurred during the disturbance period (2023-2024), characterized by strong juvenile dominance and loss of adult cohorts (?2 test of demographic structure, p < 0.001). Biomass declined by approximately 90% relative to baseline levels in 2024. Adult abundance declined substantially during disturbance years, while a juvenile recruitment pulse dominated the recovering population in 2025. However, the expected transition from medium-sized individuals in 2024 to large adults in 2025 did not occur, with only 6.7-10.5% of the predicted cohort size observed, indicating recruitment failure. Recovery trajectories differed between habitats: biomass in the swash zone rebounded to approximately 67% of baseline levels, whereas the bore zone remained strongly suppressed at 9%. Overall, the final model explained a large proportion of variance in abundance (marginal R2 = 0.75, conditional R2 = 0.79), suggesting that demographic structure and disturbance driven temperature interactions were primary drivers of population dynamics. These findings demonstrate how coastal engineering, such as beach nourishment, can trigger rapid demographic restructuring and delayed recovery through recruitment bottlenecks in surf zone macrofauna.
BY-2: A Holistic Approach to Eastern Oyster Crassostrea Virginica Restoration in the Hudson-Raritan Estuary System
Grace Schleiden (Department of Biology)
Faculty Mentors: Jason Adolf, Ph.D.
Read the Abstract—BY-2: A Holistic Approach to Eastern Oyster Crassostrea Virginica Restoration in the Hudson-Raritan Estuary System
The once abundant Eastern Oyster Crassostrea virginica population has noticeably declined in the Hudson, Raritan, and Sandy Hook estuarine system. Despite the historic decline in population size, little data has been collected on the actual population metrics in the Sandy Hook and Raritan Bay area. A huge data gap regarding oysters exists here further complicating restoration efforts. Taking a multi-faceted approach helps to propel restoration efforts while gathering knowledge needed to improve management of the species. A broad shoreline survey across the entire estuarine system will quantify local Eastern Oyster populations and locations. Reef building projects and environmental DNA experiments are more localized, being performed in the protected waters around Naval Weapons Station Earle. Environmental DNA sampling at and around oyster castle sites will be used to understand microbial, invertebrate and fish community composition, which can help determine success of the installations. By continuing restoration efforts through aquaculture and reef building, progress that has already been made can be maintained. Further knowledge on local oyster populations through research can help direct the future revitalization of the species.
BY-3: Metabarcoding the Mid-Atlantic: eDNA Insights into Elasmobranch Distribution in New Jersey
Siena Zisa (Department of Biology)
Faculty Mentors: Jason Adolf, Ph.D.
Read the Abstract—BY-3: Metabarcoding the Mid-Atlantic: eDNA Insights into Elasmobranch Distribution in New Jersey
The anthropogenic impact of climate change is affecting species distribution, especially in the Mid-Atlantic. New Jersey boasts a highly productive marine ecosystem supporting at least 336 fish species and the NJ coast serves as a critical migratory corridor. It is an important habitat for coastal sharks and other elasmobranchs. This study examined the presence and relative abundance of marine elasmobranchs using environmental DNA (eDNA) as a non-invasive monitoring tool. eDNA provides evidence of species presence, as fish shed DNA into water via mucus, scales and waste. Samples were collected quarterly over a 1 year period across seasons, from control and impact sites associated with the Orsted Ocean Wind 1 Offshore Wind Farm (OW1). Environmental parameters for each sample site were also recorded to assess drivers of fish distribution. The DNA extract was sequenced and analyzed to identify elasmobranch species present. DNA was analyzed using 12S rRNA gene metabarcoding. Geospatial analyses will be conducted to look for possible differences in elasmobranchs distribution between control and impact areas as well as seasonal differences.
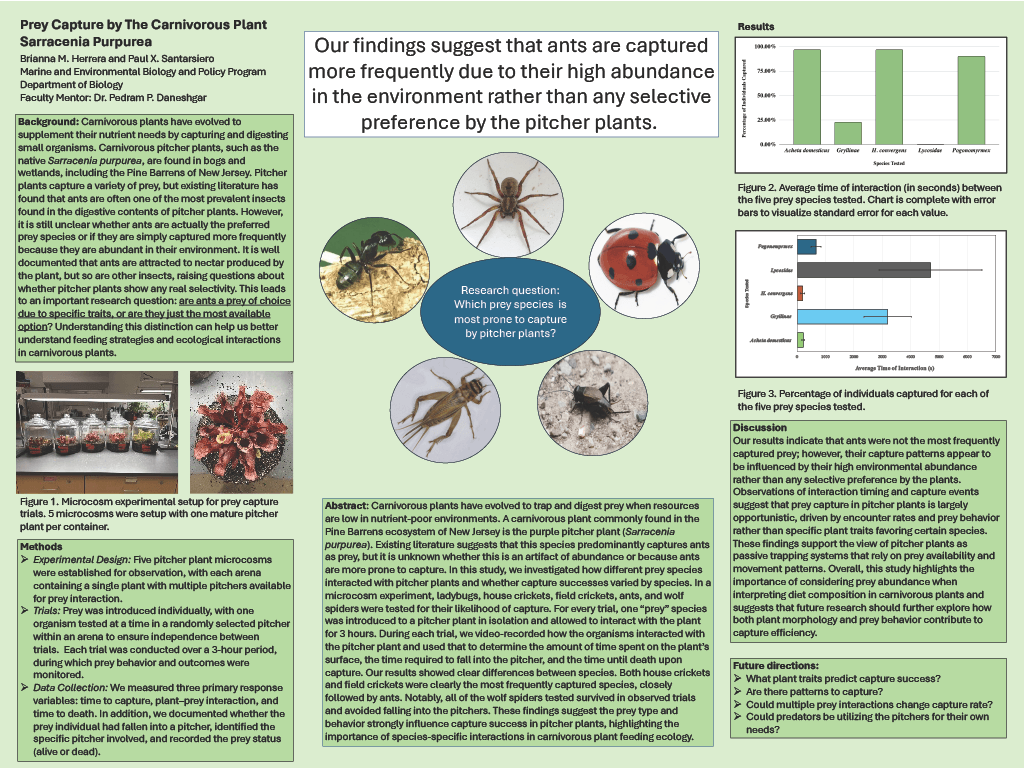
BY-4: Prey Capture by the Carnivrous Plant Sarracenia Purpurea
Brianna M. Herrera and Paul X. Santarsiero (Marine and Environmental Biology and Policy Program, Department of Biology)
Faculty Mentor: Pedram P. Daneshgar, Ph.D.
Read the Abstract—BY-4: Prey Capture by the Carnivrous Plant Sarracenia Purpurea
Carnivorous plants have evolved to trap and digest prey when resources are low in nutrient-poor environments. A carnivorous plant commonly found the pine barrens of New Jersey is the purple pitcher plant, Sarracenia purpurea. The literature suggests that these pitchers predominantly capture ants as prey, but it is unknown whether is an artifact of any abundance or because ants are prone to capture. In this study, we investigated how different prey species interacted with pitcher plants and whether capture success varied by species. In a microcosm experiment, ladybugs, house crickets, field crickets, ants, and wolf spiders were tested for their proneness to capture. One “prey” species was introduced to a pitcher plant in isolation and allow to interact with the plant for 3 hours. During each trial, we videorecorded how the organisms interacted with the pitcher plant and used that to determine the amount of time spent on the plant surface, the time required to fall into the pitcher, and the time until death after capture. Our results showed clear differences among species. Both house crickets and field crickets were most frequently captured, followed by ants, then lady beetles. Notably, almost all of wolf spiders tested survived in observed trials and avoided falling into the pitchers. These findings suggest that prey type and behavior strongly influence capture success in pitcher plants, highlighting the importance of species-specific interactions in carnivorous plant feeding ecology.
BY-5: Cypress Essential Oil Leads to JNK Activation and Apoptosis in both Fibroblasts and Fibrosarcoma Cells
Warda Chowdhury and Hermione Kissoon (Department of Biology)
Faculty Mentor: Dorothy Lobo, Ph.D.
Read the Abstract—BY-5: Cypress Essential Oil Leads to JNK Activation and Apoptosis in both Fibroblasts and Fibrosarcoma Cells
Cypress oil is an essential oil derived from evergreen coniferous trees native to Southern Europe and Western Asia. Cypress essential oil is composed of a total of 20 constituents which represent 98.1% of the oil. These include: α-pinene (48.6%), δ-3-carene (22.1%), limonene (4.6%) and α-terpinolene (4.5%) which are the main components comprising 79.8% of the oil. Some of these components have demonstrated anti-cancer properties, but little is known about their effects in fibroblasts. Previous work in our laboratory demonstrated that both HT-1080 fibrosarcoma cells and CUA-4 normal human fibroblasts displayed reduced proliferation and viability after treatment with cypress essential oil. Apoptosis was provoked, as shown by PARP cleavage, in the two cell types. The stress signaling protein JNK could be activated by cypress essential oil treatment and involved in triggering apoptosis. Both the fibrosarcoma cells and fibroblast were treated with cypress oil. Proteins were then extracted and used for western blot analysis to determine whether JNK was activated. The JNK proteins remained constant after treatment in normal fibroblasts while the level of JNK decreased after fibrosarcoma cells were treated. Activation of JNK in both cell types was still able to be identified although the level of total JNK was low in the fibrosarcoma cells. For further research, it may be possible to impede the activity of JNK with the help of chemical inhibitors or overexpression of phosphatases in order to test whether apoptosis could be reduced.
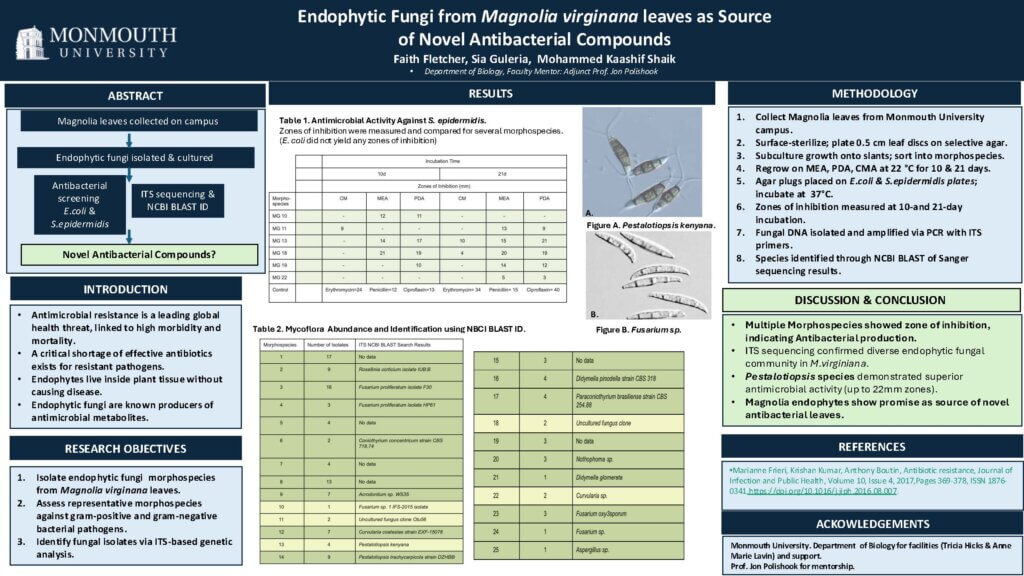
BY-6: Endophytic Fungi from Magnolia Leaves as a Source of Novel Antibacterial Compounds
Sia Guleria, Faith Fletcher, and Mohammed Kaashif Shaik (Department of Biology)
Faculty Mentor: Jon Polishook, M.S.
Read the Abstract—BY-6: Endophytic Fungi from Magnolia Leaves as a Source of Novel Antibacterial Compounds
Endophytic fungi colonize plant tissues without causing disease and have been shown to produce a wide range of bioactive secondary metabolites. As antibiotic resistance among bacterial and fungal pathogens continues to increase, endophytc fungi remain a promising source of novel antibiotic compounds. In this study, endophytic fungi were isolated from Sweetbay Magnolia (Magnolia virginiana) leaves collected from Monmouth University campus and screened for antibacterial activity. Leaf tissue, prepared as 0.5 cm discs, were surface sterilized and plated on selective agar to recover fungal species. Emerging fungal growth was subcultured to agar slants for enumeration and differentiation into morphospecies. Representative morphospecies were then grown on the production media malt extract agar plates (MEA), potato dextrose agar (PDA), and cornmeal agar plate (CMA) incubated at room temperature (22°C) for up to 21d. Agar discs of each regrown morphospecies were placed onto plates that were inoculated with a gram-positive bacterium (S. epidermatis) and a gram-negative bacterium (E. coli) and incubated overnight at 37 °C. The size of any zone of inhibition demonstrated antibacterial activity which will be compared to commercially available compounds to ascertain novelty. Genomic DNA was successfully extracted from each morphospecies and ITS PCR amplification was performed and confirmed by gel electrophoresis and Nanodrop analysis. Each morphospecies was (Sanger) sequenced by an outside vendor. Fungal species identification was ascertained through NCBI Blast searches of the resulting sequences.
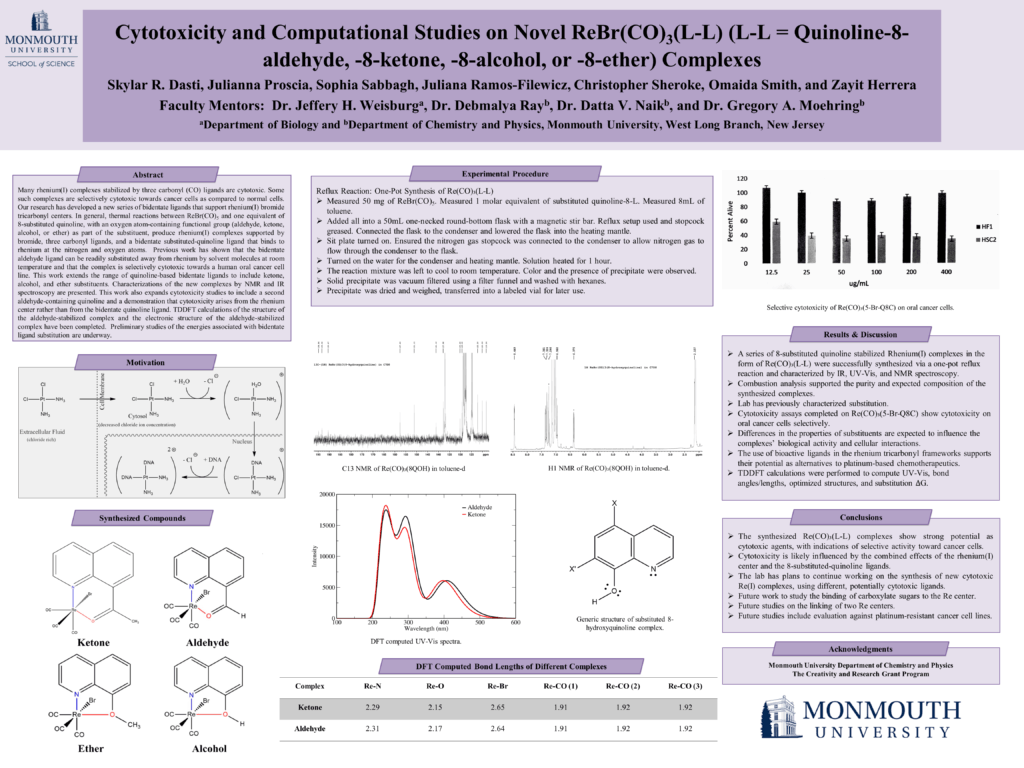
BY-7: Cytotoxicity and Computational Studies on Novel Rebr(Co)3(L-L) (L-L = Quinoline-8-Aldehyde, -8-Ketone, -8-Alcohol, or -8-Ether) Complexes
Skylar Dasti, Julianna Proscia, Sophia Sabbagh, Juliana Ramos-Filewicz, Christopher Sheroke, Omaida Smith, and Zayit Herrera (Department of Biology, Department of Chemistry and Physics)
Faculty Mentors: Jeffrey H. Weisburg, Ph.D., Debmalya Ray, Ph.D., Datta V. Naik, Ph.D., and Gregory A. Moehring, Ph.D.
Read the Abstract—BY-7: Cytotoxicity and Computational Studies on Novel Rebr(Co)3(L-L) (L-L = Quinoline-8-Aldehyde, -8-Ketone, -8-Alcohol, or -8-Ether) Complexes
Many rhenium(I) complexes stabilized by three carbonyl (CO) ligands are cytotoxic. Some such complexes are selectively cytotoxic towards cancer cells as compared to normal cells. Our research has developed a new series of bidentate ligands which support rhenium(I) bromide tricarbonyl centers. In general, thermal reactions between ReBr(CO)5 and one equivalent of 8-substituted quinoline, with an oxygen atom-containing functional group (aldehyde, ketone, alcohol, or ether) as part of the substituent, produce rhenium(I) complexes supported by bromide, three carbonyl ligands, and a bidentate substituted-quinoline ligand that binds to rhenium at the nitrogen and oxygen atoms. Previous work has shown that the bidentate aldehyde ligand can be readily substituted away from rhenium by solvent molecules at room temperature and that the complex is selectively cytotoxic towards a human oral cancer cell line. This work extends the range of quinoline-based bidentate ligands to include ketone, alcohol, and ether substituents. Characterizations of the new complexes by NMR and IR spectroscopy are presented. This work also expands cytotoxicity studies to include a second aldehyde-containing quinoline and a demonstration that cytotoxicity arises from the rhenium center rather than from the bidentate quinoline ligand. TDDFT calculations of the structure of the aldehyde-stabilized complex and the electronic structure of the aldehyde-stabilized complex have been completed. Preliminary studies of the energies associated with bidentate ligand substitution are underway.
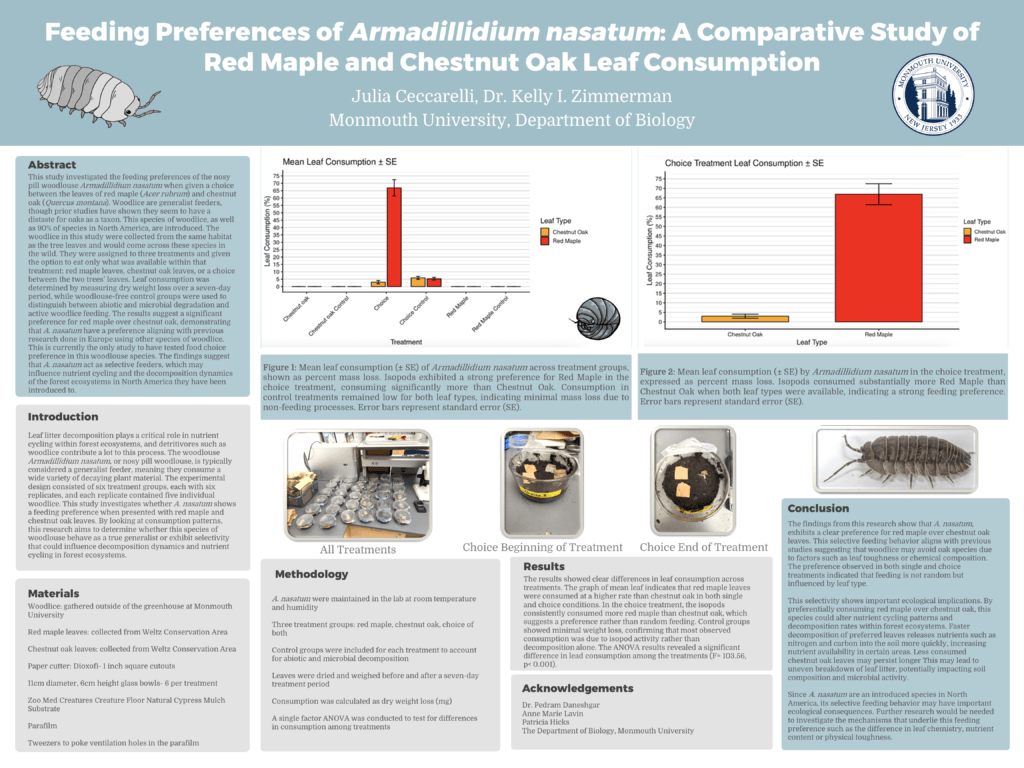
BY-8: Feeding Preferences of Armadillidium Nasatum: A Comparative Study Of Red Maple and Chestnut Oak Leaf Consumption
Julia Ceccarelli (Department of Biology)
Faculty Mentors: Kelly Zimmerman, Ph.D.
Read the Abstract—BY-8: Feeding Preferences of Armadillidium Nasatum: A Comparative Study Of Red Maple and Chestnut Oak Leaf Consumption
This study investigated the feeding preferences of the nosy pill woodlouse Armadillidium nasatum when given a choice between the leaves of red maple (Acer rubrum) and chestnut oak (Quercus montana). Woodlice are generalist feeders, though prior studies have shown they seem to have a distaste for oaks as a taxon. This species of woodlice, as well as 90% of species in North America, are introduced. The woodlice in this study were collected from the same habitat as the tree leaves and would come across these species in the wild. They were assigned to three treatments and given the option to eat only what was available within that treatment: red maple leaves, chestnut oak leaves, or a choice between the two trees’ leaves. Leaf consumption was determined by measuring dry weight loss over a seven-day period, while woodlouse-free control groups were used to distinguish between abiotic and microbial degradation and active woodlice feeding. The results suggest a significant preference for red maple over chestnut oak, demonstrating that A. nasatum have a preference aligning with previous research done in Europe using other species of woodlice. This is currently the only study to have tested food choice preference in this woodlouse species. The findings suggest that A. nasatum act as selective feeders, which may influence nutrient cycling and the decomposition dynamics of the forest ecosystems in North America they have been introduced to.
BY-9: Thermal Variability and Seasonal Migration Dynamics of Roughtail Stingray on the Jersey Coast
Angelina C. Iovino (Department of Biology)
Faculty Mentors: Keith Dunton, Ph.D., and Dane Ward, Ph.D.
Read the Abstract—BY-9: Thermal Variability and Seasonal Migration Dynamics of Roughtail Stingray on the Jersey Coast
Populations of many stingray species worldwide are in need of high conservation management efforts due to their rapidly declining populations. Seasonal variances in Sea Surface Temperature (SST) holds high importance on movement patterns of ray species. The New Jersey Department of Environmental Protection (NJDEP) established a bottom trawl survey that began in 1988, which is conducted five times throughout the year. The survey covers the entire coast of New Jersey from Sandy Hook to Cape May, utilizing a 30-meter bottom otter trawl to assess stocks. To analyze SST, we obtained data through the National Data Buoy Center (NDBC), which is a network of ocean buoys, coastal stations, and drifting platforms that continuously measure ocean and atmospheric conditions and transmit them publicly. Analysis of the NJDEP bottom trawl survey and the NDBC, helped us to identify seasonal migration patterns and abundance of Roughtail Stingray (Bathytoshia centroura) in correspondence to thermal variation along the New Jersey coast. This species showed strong patterns in abundance during the summer and early fall months (June-October) in relation to the warmer SST’s, which is displayed through the analysis of catch-per-unit-effort (CPUE). CPUE was calculated by dividing the number of captures by the number of trawls conducted, which showed the increase of ray abundance over time on the coast of New Jersey. By understanding seasonal migration in correspondence to thermal variability, we can predict future distribution patterns under climate change scenarios. This helps inform policy makers who can create strategies to protect this vulnerable species. By connecting temperature variability to movement ecology, this study explains how warming ocean temperatures may alter distribution patterns and habitat use of this species. Understanding these patterns allows scientists to better predict when and where this species will occur in.
Department of Chemsitry and Physics
CE-1: Spectroscopic Investigation of Flavonoid and Porphyrin Interactions with Human Telomeric G-Quadruplex DNA
Macklin Jugan (Department of Chemistry and Physics)
Faculty Mentors: Davis Jose, Ph.D.
Read the Abstract—CE-1: Spectroscopic Investigation of Flavonoid and Porphyrin Interactions with Human Telomeric G-Quadruplex DNA
DNA sequences rich in guanine often form structures known as G-Quadruplexes (GQs). The structures are commonly found in human telomeric sequences. Studying these molecules is compelling because GQs can regulate many biological processes, including gene expression, DNA replication, telomere maintenance, genome stability, and RNA processing. Furthermore, GQs have emerged as promising targets for the development of anticancer drugs. This study investigates the effects of selected flavonoids and a porphyrin ligand on both global and local conformational changes in GQ and duplex DNA.
Ultraviolet-visible, circular dichroism, and fluorescence spectroscopy were employed to monitor global conformational changes in GQ and double-stranded DNA upon ligand binding. To probe local structural changes, 6-methylisoxanthopterin (6MI), a fluorescent guanine base analog, was site-specifically incorporated at multiple positions within the human telomeric GQ sequence. This intrinsic fluorescent probe enabled detection of position-specific conformational perturbations within individual G-tetrads, a resolution not attainable with native guanine alone.
Three ligands were examined: TMPyP4, a well-characterized porphyrin, and two naturally occurring flavonoids, naringenin and hesperidin, which are known for their antioxidant and anticancer properties. Results demonstrated that ligand-induced structural changes are dependent on both DNA conformation and salt concentration. Furthermore, the site-specific positioning of 6MI allowed differentiation of ligand effects on distinct G-tetrads within the same molecule, revealing insights into both local and global GQ stability. These findings establish that site-specific fluorescent probes can serve as sensitive intrinsic sensors for monitoring structural and stability changes in GQs upon ligand binding. This approach provides a valuable framework for understanding drug-GQ interactions and may facilitate the rational design of targeted therapeutics for cancer and telomere-associated diseases.
CE-2: Physical Studies and Potential Cytotoxicity for a Series of Molybdenum Tetracarbonyl Complexes Stabilized by Quinoline Ligands Substituted at the 8 Position
Nicole A. Incontrera and Will H. Bradley (Department of Chemistry and Physics)
Faculty Mentors: Datta V. Naik, Ph.D., and Gregory A. Moehring, Ph.D.
Read the Abstract—CE-2: Physical Studies and Potential Cytotoxicity for a Series of Molybdenum Tetracarbonyl Complexes Stabilized by Quinoline Ligands Substituted at the 8 Position
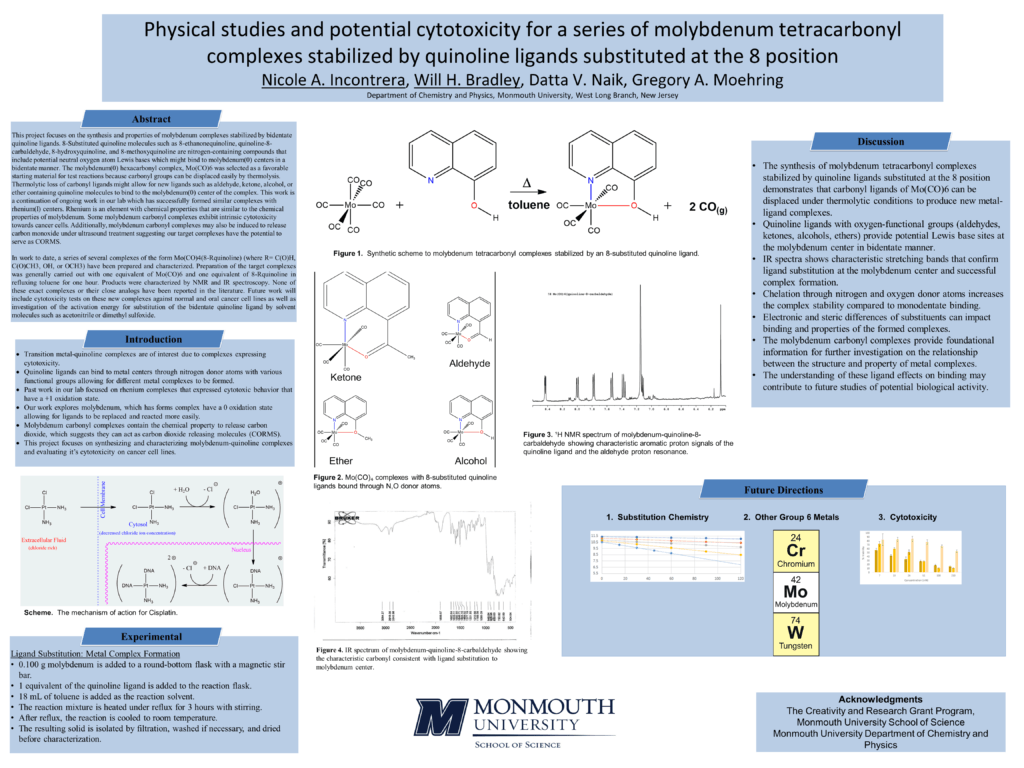
This project focuses on the synthesis and properties of molybdenum complexes stabilized by bidentate quinoline ligands. 8-Substituted quinoline molecules such as 8-ethanonequinoline, quinoline-8-carbaldehyde, 8-hydroxyquinoline, and 8-methoxyquinoline are nitrogen-containing compounds that include potential neutral oxygen atom Lewis bases which might bind to molybdenum(0) centers in a bidentate manner. The molybdenum(0) hexacarbonyl complex, Mo(CO)6 was selected as a favorable starting material for test reactions because carbonyl groups can be displaced easily by thermolysis. Thermolytic loss of carbonyl ligands might allow for new ligands such as aldehyde, ketone, alcohol, or ether containing quinoline molecules to bind to the molybdenum(0) center of the complex. This work is a continuation of ongoing work in our lab which has successfully formed similar complexes with rhenium(I) centers. Rhenium is an element with chemical properties that are similar to the chemical properties of molybdenum. Some molybdenum carbonyl complexes exhibit intrinsic cytotoxicity towards cancer cells. Additionally, molybdenum carbonyl complexes may also be induced to release carbon monoxide under ultrasound treatment suggesting our target complexes have the potential to serve as CORMS.
In work to date, a series of complexes with the form Mo(CO)4(8-Rquinoline) (where R = C(O)H, C(O)CH3, OH, or OCH3) have been prepared and characterized. Preparation of the target complexes was generally carried out with one equivalent of Mo(CO)6 and one equivalent of 8-Rquinoline in refluxing toluene for one hour. Products were characterized by NMR and IR spectroscopy. None of these exact complexes or their close analogs have been reported in the literature. Future work will include cytotoxicity tests on these new complexes against normal and oral cancer cell lines as well as investigation of the activation energy for substitution of the bidentate quinoline ligand by solvent molecules such as acetonitrile or dimethyl sulfoxide.
CE-3: Investigations of G-Quadruplex Stability Using Baby Spinach Aptamer
Ava Gaiser, Sophia Napolitano, and Erica Martin (Department of Chemistry and Physics)
Faculty Mentor: Jonathan Ouellet, Ph.D.
Read the Abstract—CE-3: Investigations of G-Quadruplex Stability Using Baby Spinach Aptamer
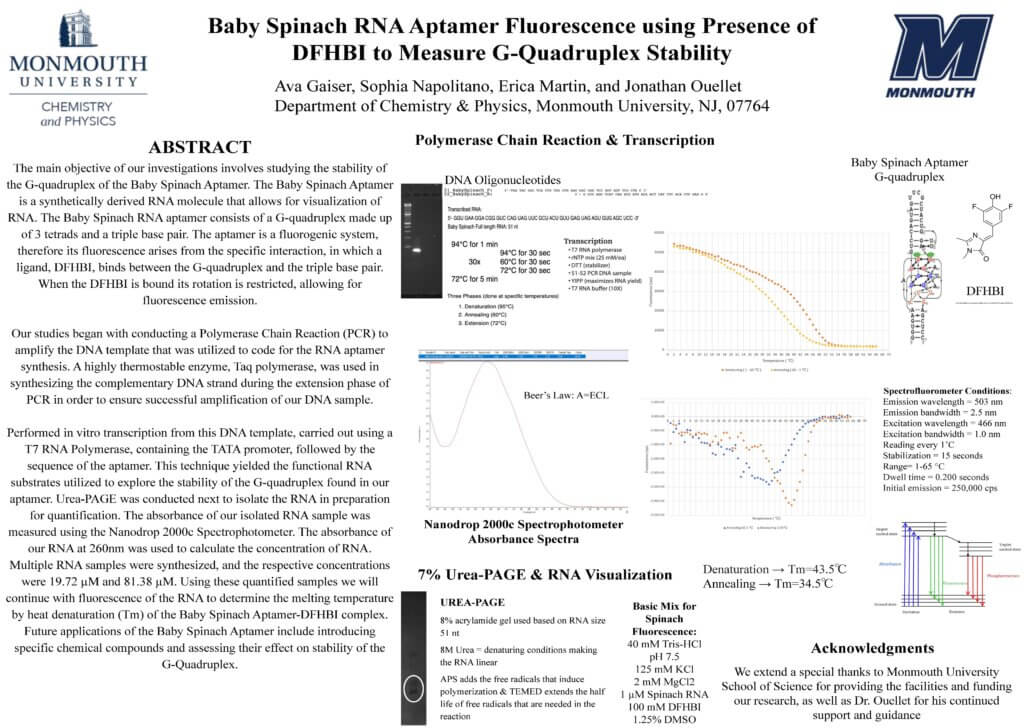
The main objective of our investigations involves studying the stability of the G-quadruplex of the Baby Spinach Aptamer. The Baby Spinach Aptamer is a synthetically derived RNA molecule that allows for visualization of RNA. The Baby Spinach RNA aptamer consists of a G-quadruplex made up of 3 tetrads and a triple base pair. The aptamer is a fluorogenic system, therefore its fluorescence arises from the specific interaction, in which a ligand, DFHBI, binds between the G-quadruplex and the triple base pair. When the DFHBI is bound its rotation is restricted, allowing for fluorescence emission.
Our studies began with conducting a Polymerase Chain Reaction (PCR) to amplify the DNA template that was utilized to code for the RNA aptamer synthesis. A highly thermostable enzyme, Taq polymerase, was used in synthesizing the complementary DNA strand during the extension phase of PCR in order to ensure successful amplification of our DNA sample.
Performed in vitro transcription from this DNA template, carried out using a T7 RNA Polymerase, containing the TATA promoter, followed by the sequence of the aptamer. This technique yielded the functional RNA substrates utilized to explore the stability of the G-quadruplex found in our aptamer. Urea-PAGE was conducted next to isolate the RNA in preparation for quantification. The absorbance of our isolated RNA sample was measured using the Nanodrop 2000c Spectrophotometer. The absorbance of our RNA at 260nm was used to calculate the concentration of RNA. Multiple RNA samples were synthesized, and the respective concentrations were 19.72 µM and 81.38 µM. Using these quantified samples we will continue with fluorescence of the RNA to determine the melting temperature by heat denaturation (Tm) of the Baby Spinach Aptamer-DFHBI complex. Future applications of the Baby Spinach Aptamer include introducing specific chemical compounds and assessing their effect on stability of the G-Quadruplex.
CE-4: Selecting for the First Glucose Aptamer
Ashley Salguero (Department of Chemistry and Physics)
Faculty Mentor: Jonathan Ouellet, Ph.D.
Read the Abstract—CE-4: Selecting for the First Glucose Aptamer
Diabetes remains a critical global health challenge, yet current management strategies are often reactive and lack the precision required for optimal care. This underscores the urgent need for advanced molecular recognition elements that enable real-time glucose monitoring and automated therapeutic responses. RNA aptamers—short nucleic acid sequences that fold into complex 3D structures—offer a promising solution due to their high stability, compact size, and ease of chemical synthesis. However, a high-affinity RNA aptamer for D-glucose has yet to be identified. This project focuses on the discovery, isolation, and characterization of D-glucosespecific RNA aptamers via Systematic Evolution of Ligands by Exponential Enrichment (SELEX). Starting with a vast library of approximately 1015 distinct sequences, the iterative SELEX method employs cycles of binding, separation, and amplification to isolate sequences with peak target affinity. While the long-term goal is to integrate these aptamers into therapeutic riboswitches, this research focuses on the essential discovery phase to overcome current limitations in molecular sensing. Successful identification of these aptamers will provide a foundational tool for synthetic biology, enabling the development of high-precision diagnostic biosensors and the engineering of autonomous, “smart” gene-regulating therapeutics for diabetes management.
CE-5: Transcription of Hammerhead Ribozyme and Spinach Aptamer to Measure with Fluorescence
Ashley Salguero (Department of Chemistry and Physics)
Faculty Mentor: Jonathan Ouellet, Ph.D.
Read the Abstract—CE-5: Transcription of Hammerhead Ribozyme and Spinach Aptamer to Measure with Fluorescence
In our investigations, we utilize in vitro for kinetic studies of hammerhead ribozyme activity. As well as exploring the folding of the Baby Spinach aptamer.
Hammerhead ribozymes, small self-cleaving RNA molecules, hold significant promise in genetic therapy due to their intrinsic catalytic RNA-cleaving activity. A comprehensive understanding of their enzymatic mechanism is crucial for optimizing therapeutic applications and elucidating the intricate RNA cleavage process. These ribozymes achieve site-specific cleavage of target RNA sequences, making them valuable tools for precise gene regulation and innovative antiviral strategies.
The Baby Spinach aptamer is a synthetically engineered RNA molecule that transforms RNA visualization in living cells. This aptamer functions as a genetically encodable, fluorogenic tag for RNA, analogous to Green Fluorescent Protein (GFP) for proteins. Its unique functionality arises from its specific interaction with the non-fluorescent ligand, 3,5-difluoro-4-hydroxybenzylidene imidazolinone (DFHBI). Upon folding into its defined three-dimensional conformation, which critically involves the formation of a G-Quadruplex, the aptamer creates a precise binding pocket. This interaction significantly enhances DFHBI’s quantum yield, inducing bright green fluorescence.
After transcription, Urea-PAGE purification and quantification by absorbance, the concentrations were 1.62µM, 3.97µM and 9.47µM for the Hammerhead Ribozyme, Hammerhead Substrate and the Baby Spinach Aptamer respectively.
The concentrations were too low for a ribozyme activity. However, a single melting temperature denaturation of the Baby Spinach Aptamer-DFHBI by fluorescence was performed, leading to a Tm of 48.0ºC.
The next steps would involve hammerhead kinetics as well as more baby spinach melting denaturation using chemical compounds that could potentially (de)stabilize the G-Quadruplex structure.
CE-6: Elucidating Binding Preferences of 5f-block Ions on CoFe2O4 Surface
Zoee Chrysanthopoulos and Amanda R. McMillen (Department of Chemistry and Physics, Department of Biology)
Faculty Mentor: Debmalya Ray, Ph.D.
Read the Abstract—CE-6: Elucidating Binding Preferences of 5f-block Ions on CoFe2O4 Surface
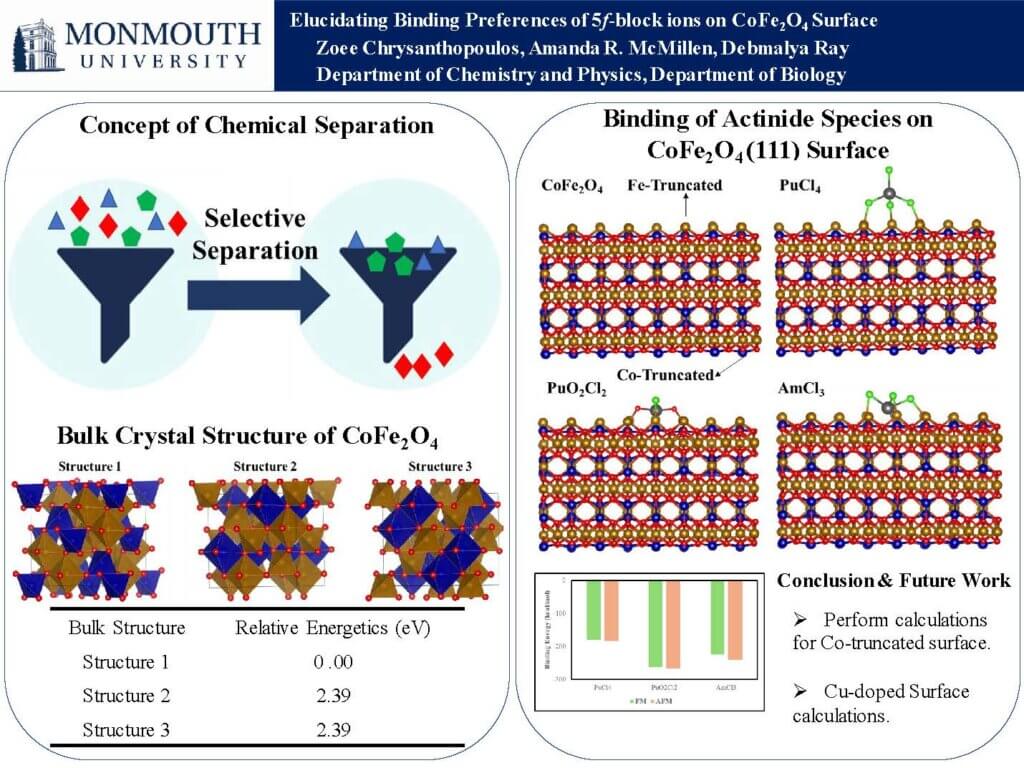
Due to its low greenhouse gas emissions and affordability, nuclear energy has emerged as a clean and efficient power source, addressing the escalating global energy demand while mitigating the adverse effects of climate change. A critical aspect of ensuring the sustainable and responsible use of nuclear energy is the effective management of nuclear waste. This involves the safe disposal of spent nuclear fuel and the recovery of valuable fissile elements like plutonium and americium. In this project, we explored the performance of cobalt ferrite crystal structure to sequester Tri (Am3+), Tetra (Pu4+), and Hexavalent (PuO22+) actinides from aqueous acidic solutions. We first determined the most stable structure of the cobalt ferrite by substituting cobalt on different sites of Fe3O4 crystal structure. Further we evaluated the binding of the actinide ions on the 111 crystallographic surfaces of CoFe2O4. We used density functional theory based computational methods to explore the structure and binding relationship of actinide ions on the CoFe2O4 structure. Our computational result shows that the selectivity of binding these ions are Pu(VI) > Pu(IV) > Am(III). We believe, our computational work will pave the path for future experimental studies on use of Cobalt Ferrite for radioactive waste water remediation and how to reprocess valuable f-block elements for nuclear energy applications.
CE-7: Computational Understanding of Host-Guest Interaction of Supramolecular Complex with Biologically Active Molecules
Sherlyn Garcia and Lauren Manrique (Department of Chemistry and Physics, Department of Biology)
Faculty Mentor: Debmalya Ray, Ph.D.
Read the Abstract—CE-7: Computational Understanding of Host-Guest Interaction of Supramolecular Complex with Biologically Active Molecules
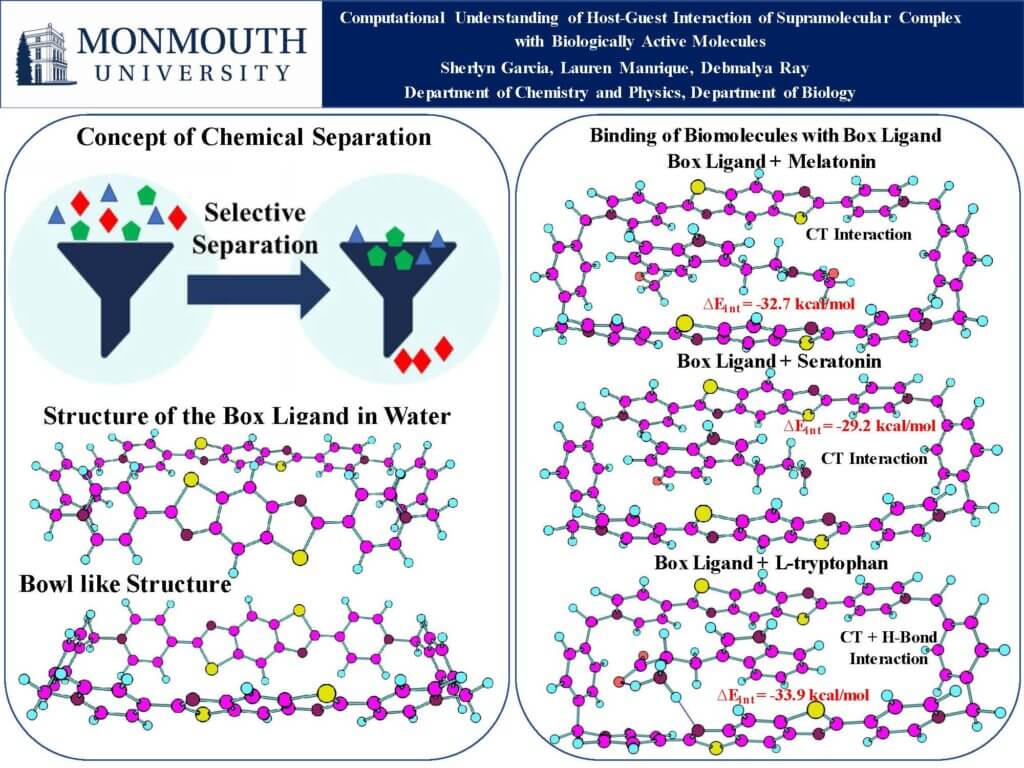
The separation of L-tryptophan, serotonin and melatonin is crucial in both biological research and clinical diagnostics because they serve distinct roles in human physiology while sharing a tightly linked metabolic pathway. While tryptophan is the essential amino acid precursor, serotonin acts as a neurotransmitter regulating mood and cognition, and melatonin is the hormone that regulates the sleep-wake cycle. In this work we designed supramolecular complexes that can selectively bind melatonin, serotonin and L-tryptophan in water solvent. Our study highlights how charge transfer plays a role in forming host-guest interactions in these complexes
CE-8: Computational Understanding of Interaction of Gold (Au13) Nanocluster with CBD/CBN for Medical Drug Delivery Applications
Alex Pante and Brianna Klohn (Department of Chemistry and Physics, Department of Biology)
Faculty Mentor: Debmalya Ray, Ph.D.
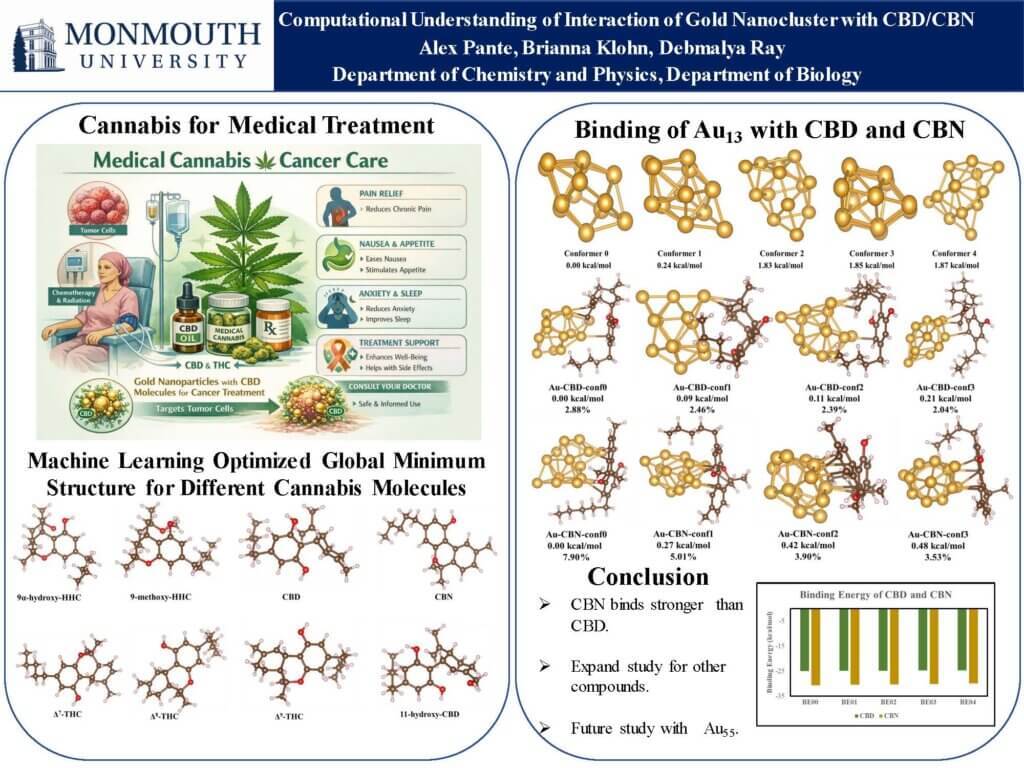
Read the Abstract—CE-8: Computational Understanding of Interaction of Gold (Au13) Nanocluster with CBD/CBN for Medical Drug Delivery Applications
Metallic nanocarriers, including gold, silver, and iron oxide nanoparticles and nanoclusters, exhibit unique surface properties that enhance the anticancer efficacy of drugs. Further, experimental work showed promise of using Cannabis for various medical treatments. Although Cannabis class of molecules showed some promise for medical treatments there are limited research related to efficient delivery or combinatory therapy of cannabis delivery with metal nanoclusters. Thus, in this work we explored the interaction of two different cannabis molecule CBD and CBN with Au13 nanoclusters and computed the thermodynamics of these interactions. Our study shows promise of using gold nanoclusters for CBD and CBN delivery which is relevant to medical drug delivery applications.
CE-9: Surface Interactions of Goethite with PET Microplastics: Insights from FT-IR & XPS Characterization
Isabella R. Hallingse, Thomas P. Smith, and Jacob W. George (Department of Chemistry and Physics)
Faculty Mentor: T. Tongesayi, Ph.D.
Read the Abstract—CE-9: Surface Interactions of Goethite with PET Microplastics: Insights from FT-IR & XPS Characterization
This study examined how goethite (GT) alters the surface chemistry of thermoplastic‑bead microplastics (TPB‑MPs) and polyethylene terephthalate microplastics (PET‑MPs) under simulated natural conditions. GT was selected due to its environmental abundance and its known influence on particle behavior and pollutant interactions. Microplastic surfaces were analyzed before and after GT exposure using X‑ray photoelectron spectroscopy (XPS), and prior Fourier‑transform infrared (FT‑IR) analysis confirmed that TPB‑MPs are PET‑based materials with similar chemical composition. XPS characterization revealed changes in elemental composition and electron binding energies associated with GT adsorption. GT treatment decreased surface carbon and increased oxygen content more strongly in PET‑MPs than in TPB‑MPs. PET‑MPs exhibited evidence of stronger charge transfer and hydrogen bonding with GT, whereas TPB‑MP interactions were weaker and dominated by van der Waals forces. Variations in peak intensities indicated enhanced C–O and O–C═O bonds and masking of C–C/C═C bonds in PET‑MPs, while TPB‑MPs showed fewer chemical modifications. Shifts in the Fe2p doublet further suggested chemical changes associated with GT adsorption. Overall, the findings demonstrate that GT significantly modifies the surface chemistry of PET‑MPs, enhancing their environmental reactivity and potential for transformation, whereas TPB‑MPs exhibit weaker interactions and fewer surface alterations. These results highlight GT’s role in mediating MP–mineral interactions and suggest that future research should investigate oxidative pathways and microbial responses to mineral‑coated microplastics.
Department of Computer Science and Software Engineering
CSSE-1: Quantum Machine Learning for Phishing Email Detection Using Quantum Support Vector Machines
Sophia Ramirez Velandia (Department of Computer Science and Software Engineering)
Faculty Mentor: Brian Callahan, Ph.D.
Read the Abstract—CSSE-1: Quantum Machine Learning for Phishing Email Detection Using Quantum Support Vector Machines
Phishing emails remain one of the most common and costly cybersecurity threats, responsible for a significant percentage of successful data breaches worldwide. Attackers frequently design phishing messages that closely imitate legitimate communication from trusted institutions such as banks, universities, and online services. Because these messages often resemble normal emails in tone, structure, and formatting, they can be difficult to detect using traditional filtering techniques. While classical machine learning models have improved phishing detection in recent years, they often struggle when malicious messages closely resemble legitimate communication. This research investigates whether quantum machine learning techniques can improve phishing detection performance by identifying patterns that may not be easily separable within classical feature spaces.
To conduct this study, several publicly available email datasets containing both phishing and benign messages were collected and processed. A preprocessing pipeline was developed to convert raw email text into numerical feature representations suitable for machine learning analysis. These features include keyword frequency, structural email metadata, and behavioral indicators commonly associated with phishing attacks, such as suspicious link frequency, urgency-based language, abnormal punctuation patterns, and message length characteristics. The data was then cleaned and standardized to ensure consistent input for machine learning models.
The extracted features were encoded into a quantum feature space using a parameterized quantum circuit designed to represent classical data within a higher-dimensional Hilbert space. Similarity between samples was calculated using a fidelity-based quantum kernel, which measures the overlap between quantum states. A Quantum Support Vector Machine (QSVM) classifier was trained using this kernel and compared against a classical Support Vector Machine baseline. Experimental results show that the quantum classifier achieved approximately 73.3% classification accuracy, outperforming the classical SVM baseline of approximately 60% accuracy. Additional evaluation using five-fold stratified cross-validation confirmed the stability of the quantum model, with an average accuracy of approximately 67%.
An important challenge in quantum machine learning is the presence of noise in current quantum hardware, including decoherence, gate errors, and measurement uncertainty. Kernel-based quantum approaches may offer some resilience to noise because similarity between samples is calculated through quantum state overlap rather than relying solely on individual gate operations. While the experiments in this study were conducted primarily using a noiseless simulator, ongoing work is exploring execution on real quantum hardware to evaluate how the model performs under realistic noise conditions and to better understand its robustness under hardware-induced errors.
CSSE-2: Cyberbeangames: The Gamification of Cybersecurity Education
Lynda Levy (Department of Computer Science and Software Engineering)
Faculty Mentor: Weihao Qu, Ph.D., Brian Callahan, Ph.D.
Read the Abstract—CSSE-2: Cyberbeangames: The Gamification of Cybersecurity Education
As digital threats continue to evolve, engagement in cybersecurity education has seen a marked decline, traditional methods such as lengthy instructional videos followed by perfunctory quizzes have proven insufficient in fostering meaningful awareness of online safety. This research addresses that gap by offering a framework for more effective cybersecurity education, with particular focus on internet safety and phishing scam prevention, applicable to both academic and professional settings. Utilizing CyberBeanGames.com, a website created by students and professors at Monmouth University, we can empower students to design and develop educational games. This approach serves a dual purpose: reinforcing the creators’ own understanding of the subject matter while simultaneously providing an engaging and accessible learning experience for all participants.
Data was collected from 11 participants between February and March 2026. 100% of participants gave the platform a rating of 5 out of 5, demonstrating user satisfaction with the platform. Post‐game quiz scores were consistently higher than pre‐game scores among all participants who completed a game, with 5 out of 6 participants who completed a post‐game quiz scoring perfectly. Despite most users coming from a CS/IT background, participants with no prior training still performed well post‐game, suggesting the games are effective regardless of background.
Traditional cybersecurity education methods have proven insufficient in producing lasting awareness among students and professionals. CyberBeanGames.com, developed by students and faculty at Monmouth University, offers a more engaging alternative. With all participants rating the platform 5 out of 5 and post-game scores consistently outperforming pre-game scores, the findings confirm that game-based learning drives stronger knowledge retention while empowering student creators in the process.

CSSE-3: The Data Will Flow: Measuring How Embodied Performance Models Environmental Data
Brooke Tortorelli (Department of Computer Science and Software Engineering)
Faculty Mentor: Jiacun Wang, Ph.D.
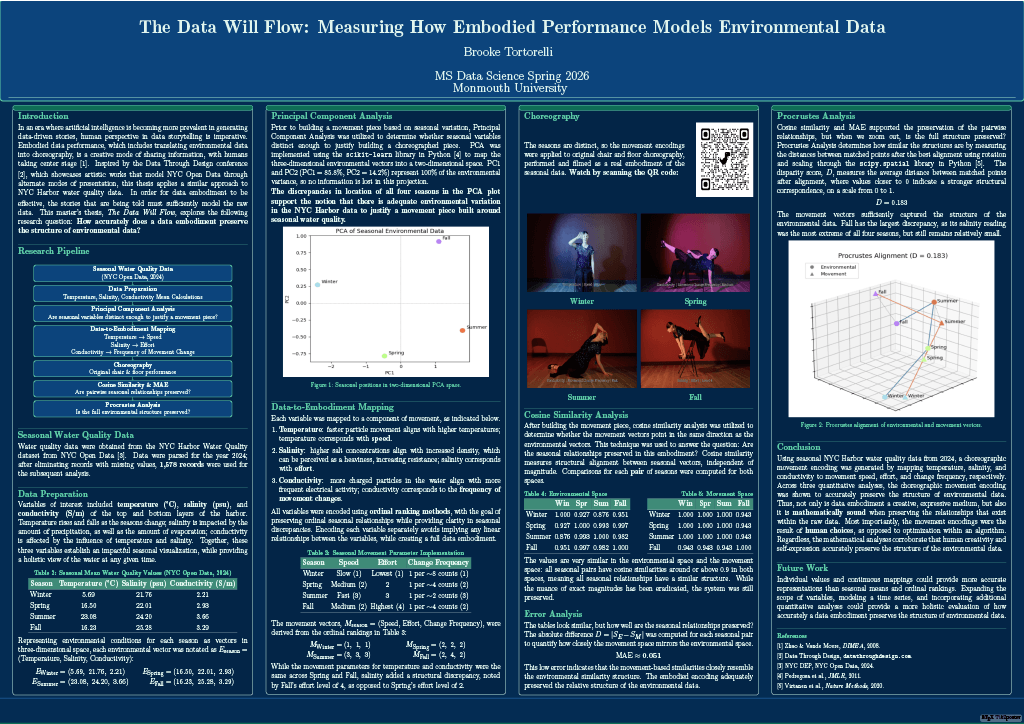
Read the Abstract—CSSE-3: The Data Will Flow: Measuring How Embodied Performance Models Environmental Data
In an era where artificial intelligence is becoming more prevalent in generating data-driven stories, human perspective in data storytelling is imperative. Embodied data performance, which includes translating environmental data into choreography, is a creative mode of sharing information, with humans taking center stage. In order for data embodiment to be effective, the stories that are being told must sufficiently model the raw data. This thesis poses the question: how accurately does a data embodiment preserve the structure of environmental data? To explore this, water quality data from NYC Harbor were obtained from NYC Open Data. Data were filtered to 1,578 complete records from 2024 and divided into four seasons. Three variables, temperature, salinity, and conductivity, were mapped to choreographic movement parameters: movement speed, effort, and frequency of change, respectively. These mappings were applied to original chair and floor choreography, performed and filmed as a real embodiment of the seasonal data. To evaluate how well the movement encoding preserved the structure of the environmental data, four quantitative analyses were performed, including Principal Component Analysis, cosine similarity analysis, Mean Absolute Error analysis, and Procrustes Analysis. The results indicated that there were distinct qualities within the variables for each season, relationships between seasons were maintained, and structural differences between environmental and movement spaces were minimal in two- and three-dimensional spaces. Not only is data embodiment a creative, expressive medium, but also it is mathematically sound when preserving the relationships that exist within the raw data. Most importantly, the movement encodings were the result of human choices, as opposed to optimization within an algorithm.
CSSE-4: Decked Out: A Physics-Based Platform Fighting Game
Roman Dattilo, Mike Montulet, Camryn Moschitta, Mike Cozzolino, and Justin Veltri (Department of Computer Science and Software Engineering)
Faculty Mentor: Cui Yu, Ph.D.
Read the Abstract—CSSE-4: Decked Out: A Physics-Based Platform Fighting Game
Decked Out is a multiplayer platform fighting game developed in Unity, inspired by games such as Super Smash Bros. The goal of the project was to create an enjoyable and unique experience that players can share with friends. Players select their character by drawing a card, with a rosterof personified creatures, each featuring distinct playstyles and abilities.
A core focus of the project was designing smooth, responsive gameplay. This included implementing player movement using Rigidbody linear velocity, hit detection through collision-based triggers, and synchronizing animations with attacks using keyframe events. Another key feature is local two-player multiplayer, where the game detects connected controllers and assigns them to player slots automatically.
The project also emphasized creating original characters. The team developed character artwork, 3D models, and move sets for each fighter. The current roster includes five characters: Winston Woodchuck, Vera Voidwell, Brutus Boxshell, Jolly Archie, and Otis O’Mud.
The result is a fully playable video game with a functional user interface and main menu. Core gameplay has been completed, and development is ongoing as additional characters are animated and integrated into the roster.
Department of Mathematics
MA-1: Prey Susceptibility to Capture by Purple Pitcher Plants (Sarracenia purpurea)
William Ahlfeld, John Caufield, and Michael Yax (Department of Mathematics)
Faculty Mentor: Pedram Daneshgar, Ph.D. (Department of Biology), and Richard Bastian, Ph.D. (Department of Mathematics)
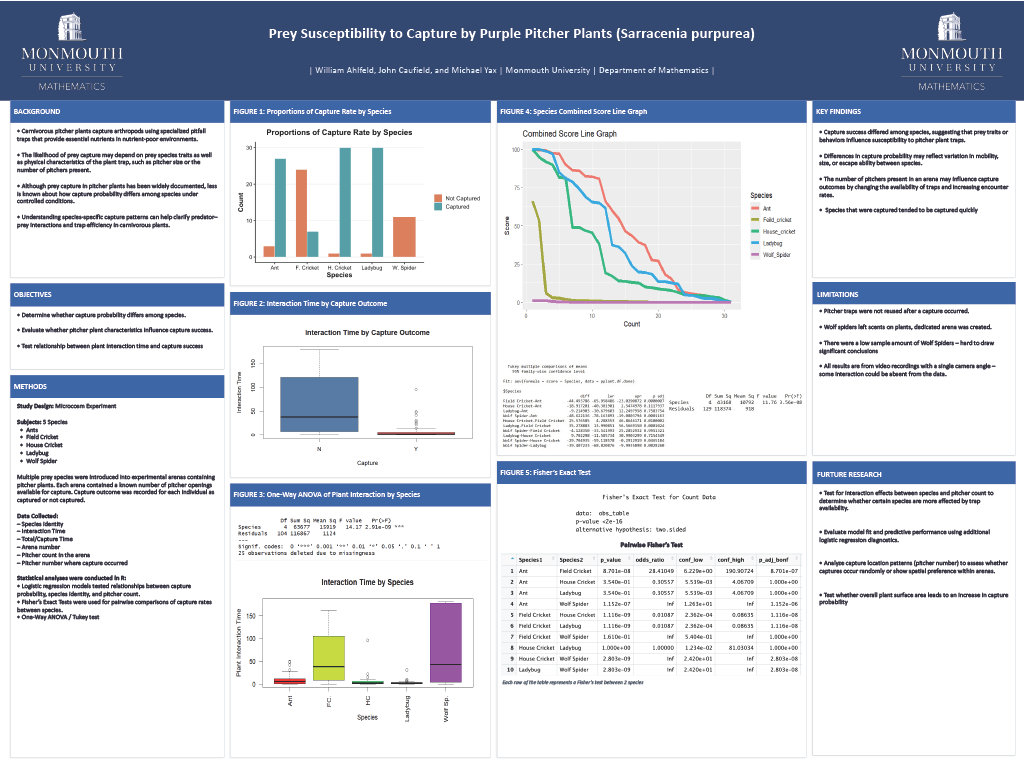
Read the Abstract—MA-1: Prey Susceptibility to Capture by Purple Pitcher Plants (Sarracenia purpurea)
Previous diet analysis of the carnivorous purple pitcher plant has suggested that ants are the primary prey captured. Is this because ants are easy to capture? Is this because ants are more abundant than other prey species in the pitcher habitat? We experimentally examined the capture rate of five different prey species (ants, house crickets, field crickets, ladybugs, and wolf spiders) by pitcher plants in a microcosm experiment. Individual prey were placed with a pitcher plant in isolation and video recorded for 3 hours so that their interaction could be observed. Interactions were timed, and mortality (if any) was recorded, including time of interaction with plant, time to capture and time to die. Pitcher morphology (e.g., volume, space, and number of traps) was also measured as an independent variable. Five arenas were used, each containing a single purple pitcher plant (Sarracenia purpurea), with arenas rotated to minimize confounding variables such as spider scent trails. The goal is to determine if ants are primarily captured due to susceptibility or environmental abundance, and which species is most prone to capture overall. Preliminary statistical analysis will compare capture rates, interaction times, and mortality across species using variables like arena number, plant traits, and interaction times. Results will provide insights into pitcher plant ecology, informing conservation efforts amid growing popularity of carnivorous plants in the horticultural industry.
MA-2: TPLO Post Operative Infection Rate Analysis
Royce Amburg, Brandon Bloch, and Matthew Muller (Department of Mathematics)
Faculty Mentors: Richard Bastian, Ph.D. (Department of Mathematics) and Dr. Sara Mansuri (Red Bank Veterinary Hospital)
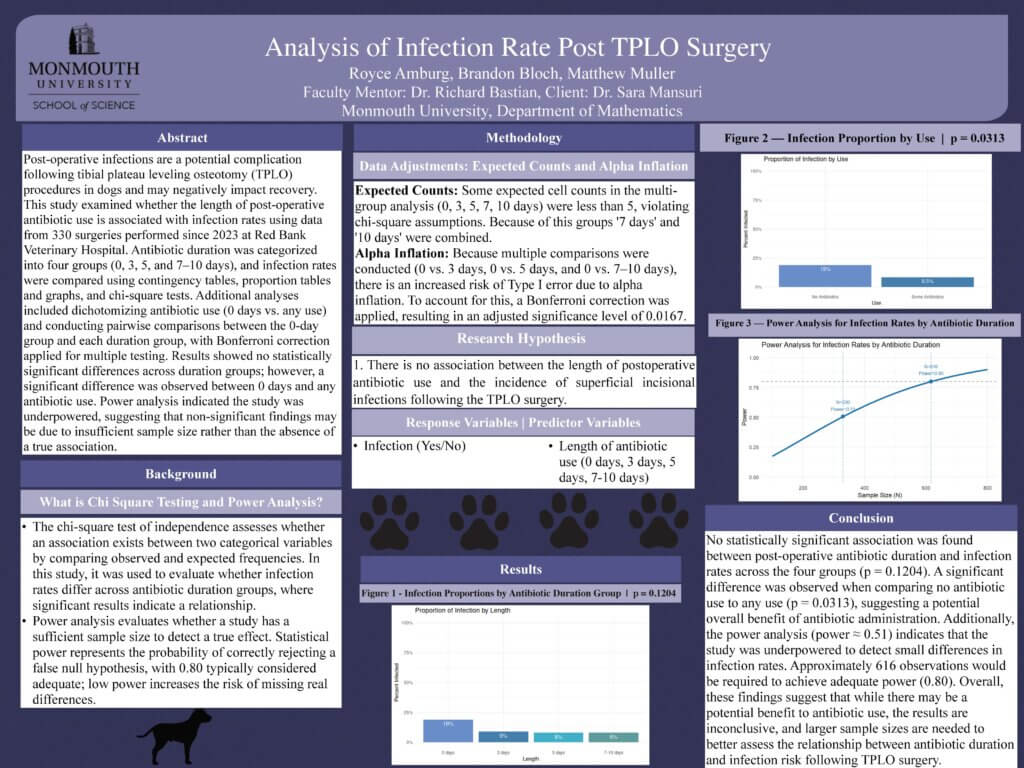
Read the Abstract—MA-2: TPLO Post Operative Infection Rate Analysis
Post operative infections are a potential complication following tibial plateau leveling osteotomy (TPLO) procedures in dogs and can negatively impact recovery. This study examines whether the length of post-operative antibiotic use is associated with different infection rates following TPLO surgery. Data from 330 TPLO surgeries performed on dogs since 2023 from Red Bank Veterinary Hospital in New Jersey were analyzed. The primary analysis compared infection rates across groups of dogs receiving different durations of post-operative antibiotics. Antibiotic use or ‘length’ was categorized into four groups based on treatment: 0 days, 3 days, 5 days and 7-10 days. Infection rates were compared across these groups using contingency tables, proportion tables and chi-squared tests to determine whether meaningful differences were found among groups of treatment durations. Additionally, antibiotic use was dichotomized (0 days vs. any use) and, pairwise comparisons between the 0-day group and each antibiotic duration group were conducted, with Bonferroni correction applied to adjust for multiple testing. A power analysis was done to evaluate whether the sample size of 330 dogs provided sufficient statistical power to detect differences in infection rates across antibiotic length groups. The results showed that there were no statistically significant differences in infection rates across antibiotic duration groups however there was a statistically significant result comparing the groups of 0 days and any use of antibiotics. The power analysis indicated that the study was underpowered (power ≈ 0.51), suggesting that the non-significant findings may be attributable to insufficient sample size rather than the absence of a true association.
MA-3: Factors Contributing to the Rupture of Splenic Masses in Dogs
Aleah Lazar and Vincent Macri (Department of Mathematics)
Faculty Mentors: Richard Bastian, Ph.D. (Department of Mathematics) and Dr. Kevin Cali (Garden State Veterinary Specialists)
Read the Abstract—MA-3: Factors Contributing to the Rupture of Splenic Masses in Dogs
Splenic masses are incredibly common in dogs and are highly significant due to the high likelihood of malignancy and risk of rupture. Without emergency surgery, a ruptured mass is often life threatening. Understanding the factors that relate to higher incidence of rupture is incredibly important when determining if a dog needs surgery. This study focuses on the number of masses a dog has and their location within the spleen. Retrospective veterinary records of 726 dogs who had undergone surgery were analyzed to determine patterns in rupture incidence according to the aforementioned factors. Preliminary findings suggest that dogs with multiple masses (4+) may have an increased risk of rupture. Masses in the head of the spleen were also seen to rupture more than masses in the body or tail of the spleen. Furthermore, multiple masses located in one location in the spleen may also be associated with an increased risk of rupture. The results of this study are important in ensuring both doctor and patient have all the information necessary while determining if surgery is worth it.
